 In this tutorial, we will be using the x-ray crystal structure of the human RAMP1 extracellular domain (PDBID: 2YX8). What it is that you are trying to model and choose a PDB accordingly. Several studies solve mutated proteins, or solve it in complex with small molecules. (or absence) of inhibitors, cofactors, or substrates. Consider the resolution, the sequence, and the presence When selecting a PDB for your protein of interest, do not select the first PDB that you see. Each PDB has a unique code, called a PDBID, that allows it to be found and referenced quickly. A PDB is a file format that contains 3D coordinates for a biomolecule that were determined using an experimental method suchĪs X-ray crystallography, cryo-EM, or NMR spectroscopy. The Protein Data Bank (PDB), found at, is a central repository of experimentallyĭetermined structures. Download a pdb file from the PDB Databank. To see the path to your working directory, you can use Make a directory for your ~]$ mkdir tutorial ~]$ cd tutorial/ 
For help learning the command line in Unix, see:ġ. Use VMD Visualization Software to Examine the Protein StructureĪmberTools is used through Unix command line, which is accessed via a This tutorial requires AmberTools and VMD be properly installed More experienced users who are building a protein system for the first It is targeted toward beginners however it may also be useful to This tutorial will go over how to build a protein system using AmberTools. Use LEaP to build a protein system in explicit solvent.Evaluate and analyze the protein structure to be imported into Amber.Use VMD to examine a protein structure.Use LEaP to Build a Protein System in Explicit Solvent Evaluate and Analyze the Protein Structure to be Imported into AmberĬ) Experimental methods noted in the paper associated with the PDBĭ) Solvent molecules or crystallization bufferĮ) Missing electron density (amino acids) Use VMD Visualize Software to Examine a Protein Structure 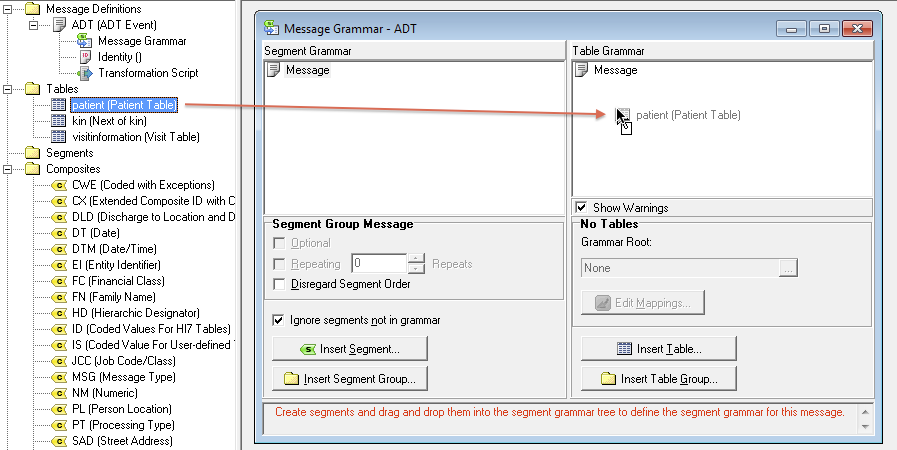
This nearly replicates Qutemol-like rendering, but you’ll want to play with the colors that you use.1.5 Building Protein Systems in Explicit Solventġfrom Bill Miller III's lab, Truman State University, 2Stony Brook University Table of Contents I.Introduction To give a soft appearance that emphasizes depth, we can use Tachyon with a diffuse material. Now, the result of our rendering can differ greatly depending on which renderer we use. As for the rest of the settings, see the previous tutorial for turning on ambient occlusion, setting the material to diffuse, and Drawing Method as CPK with Sphere Scale/Bond Scale 1.6 and 1.0. You can choose an alternative option by going to Graphics>Colors and choosing the Color Scale tab (I’ll actually be using red/green/blue today with an offset of 0.30 to make the colors lighter). The color gradient by default is red/white/blue. Now we can go back to the Draw style tab and choose coloring method as Timestep.  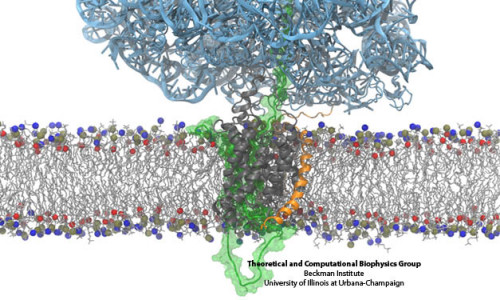
Since there are 15 frames numbered from 0 to 14 and we want to see all of them, we’ll write 0:14 in the Draw Multiple Frames dialog box. Go to Graphics>Representations and select the Trajectory tab. In VMD, we can also overlay all of the different structures from our trajectory at the same time. Go to Extensions>Analysis>RMSD Trajectory Tool and you’ll see a window that looks like this: You’ll notice that the different structures are not aligned in any way. You can look at the different coordinates by sliding back and forth the bar on the bottom of the VMD main window. When you open this file, you’ll notice that there is more than one set of coordinates in it. Start by opening the PPant-Anim.xyz file in VMD. Now, there are a few details of VMD we didn’t get to last time that are useful for getting more descriptive figures from your coordinate files with VMD. Revisiting VMD: Trajectories, Tachyon and beyond. Nevertheless, all you really need to know is that it is the example molecule we’ll work with today, and you can download the coordinates we’ll use for the visualization here and here.ġ. This prosthetic tether forms a thioester linkage to amino acids and delivers them to the active sites of many different enzymes found most commonly in natural product biosynthetic pathways. We’ll use a model for the phosphopantetheine tether of acyl carrier proteins. As a continuation of the previous tutorial, I will show you a few more things you can do with VMD along with some tricks for PyMOL. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
